The Group is on lockdown but work continues:
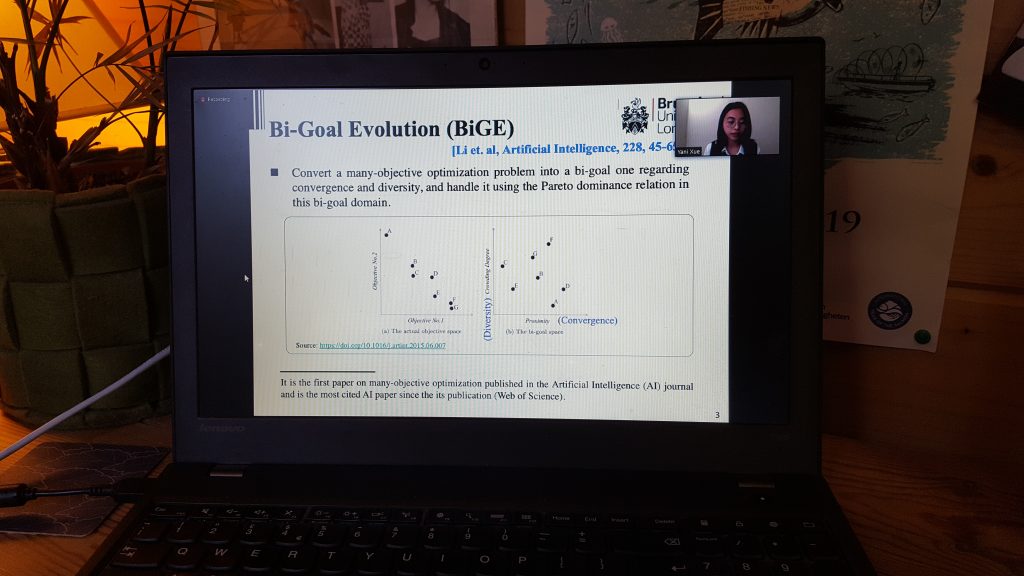
Seminars are continuing online
Some staff are volunteering their hardware and data analysis skills at Hillingdon hospital
A collaboration with the MHRA and CPRD will result in synthetic primary care data being made available for Covid19 research.
Biraja Ghoshal has been exploring his methods for identifying Covid19 from lung images: https://arxiv.org/abs/2003.10769